
Tl2, a new locus resembling Tl in its action
Berdnikov, V.A. and Gorel, F.L. Inst. of Cytology and Genetics, Russian Academy of Sciences
Novosibirsk, Russia
Pea possesses the most complex compound leaves of all genetically studied plant species. Its unipinnate leaf is composed of a pair of basal stipules, one or more pairs of proximal leaflets, one or more pairs of distal tendrils, and a single terminal tendril. The gene Tendril-less (tl), together with Afila (af) and Unifoliata (uni), belongs to a group of genes able to drastically alter the leaf architecture. The mutation tlw, appears to replace the tendrils of the distal region of the leaf by normal oval leaflets. The Tl gene displays incomplete dominance so that heterozygotes tlw/+ in the distal region of the leaf possess pinnae morphologically intermediate between tendrils and leaflets, we call them ‘Flat Tendrils‘ and designate the leaf phenotype of the heterozygotes tlw/+ as FT. It should be noted that the wild-type allele Tl not only represses lamina formation in the pinnae of the distal part of the leaf but also contributes to branching potential of the rachis, this effect being especially evident in a af/af background.
Examination of the M2 generation produced by treating the line SG (Tl, R) with g-rays (7000R), revealed a plant with tendrils converted to very narrow leaflets (Fig. 1). Its phenotype resembled FT, exhibited by heterozygotes tlw/+, suggesting that it was a new mutation at the Tl locus. We crossed this plant with the line WL1238 (tlw r). Of seven F1 plants examined, five had the phenotype FT and two had the phenotype tendril-less (tl). The appearance of these phenotypes is consistent with the hypothesis that the mutant plant with flat tendrils was heterozygous tl/+. The F1 plants with the phenotype tl produced 77 seeds, of which 29 were wrinkled and 48 round, indicating that segregation in the region around R was relatively normal. However, when we grew out the F2, two unexpected phenotypic classes with respect to tendril shape were obtained. The F2 contained three phenotypic classes: tl (42 plants), FT (26 plants) and N (9 plants with wild-type leaves). All of the 29 plants grown from the wrinkled seeds had the phenotype tl. This result was not surprising considering the tight linkage between the loci R and Tl. The segregation pattern indicated that the mutant in the line SG affected a hitherto unknown gene conditioning development of flat tendrils (FT) upon its inactivation. We designated this gene as tendril-less2 (tl2).

Fig. 1. Phenotypes of the tl2/Tl2 heterozygotes of the line Flat-1: left, young leaf; center, mature leaf; right, distal portion of a compound leaf.
A plant with the phenotype FT, homozygous for R and supposedly heterozygous for tl2, was crossed with the line Sparkle (r, a, Tl) and for three generations a single most vigorous plant
with the phenotype FT
was selected. Self-pollination
of these plants never produced plants with the phenotype tl, while
wild-type and FT phenotypes
were present in approximately equal proportions, the latter slightly
predominating. Such segregation pattern suggests that in the case of the tl2
gene, the FT phenotype reflects heterozygosity
at Tl2. The
absence of the tl phenotype
suggests lethality of the homozygotes tl2/tl2. The explanation for the observed FT:N ratio of close to 1:1 instead of 2:1 may reside in reduced transmission
of the lethal through the germ line. Two F4 plants with the FT
phenotype
became progenitors of the lines Flat-1 (a,
tl2/+, R) and Flat-2 (A,
tl2/+, R). We made an extensive cross between the line Flat-1 (♀) and the line DSP (a, Tl) (♂)
derived from cv. Dark Skinned Princess. The success of the cross was confirmed
by electrophoretic analysis of the cotyledon protein SCA.
The F1 produced 112 seeds and 97 F2 plants, of which 48 had the phenotype FT and 49 the wild-type phenotype. The striking equality in the numbers of both
phenotypic classes showed that transmission of tl2 through female germ line did not
differ from the wild-type allele Tl2. We conclude that the deviation of
the phenotype ratio in the progeny produced by self pollination is due to a
decreased transfer of tl2 through
the male germ line. It should be noted that all FT plants had fully fertile pollen implying that no translocation was involved.

Lethality and the difference in gamete competitive ability precludes standard analysis of recombination relationships in the F2. However, if tl2 is tightly linked to some marker, the proportion of plants with this marker should be strongly biased in the classes FT and N. An example of such an analysis is given in Table 1. If the loci Tl2 and R were tightly linked, the plants grown from the wrinkled seeds would mostly have the wild-type phenotype; however, in the experiment we did not observe a noticeable deviation in the ratio FT:N in classes r and R. We tested the loci Curl, Wlo, Uni. I, Af, and Cri in the same manner and found no evidence of linkage with Tl2.
 To check more loci, we crossed the line Flat-1
with our line HT1 (A,
Tl2, gp)
homozygous for the Hammarlund translocation (Table 2). (This
line was described in ref. 4). Analysis of 119 F2
plants revealed an obvious linkage of Tl2 with the translocation breakpoint
and the locus Gp residing near the breakpoint in
chromosome V
(here we use the linkage group numbers for desig-nation of the corresponding chromosomes). The major-ity (54 of 65) of the
structural heterozygotes, registered by pollen semi-sterility had the phenotype FT, that is, were hetero-zygotes tl2/+. The majority of fully fertile
plants (44 of 54) had wild-type tendrils. One can conclude that Tl2 resides near the translocation
breakpoint. In the line HT1 the longer interchange chromosome bears the allele gp near the T-point. Thirty-two
of 35 F2 plants
with the phenotype gp had
normal tendrils and only three had flat tendrils. These data provide evidence in favor of tl2 being located on a non-translocated
chromosome, either V or II.
To check more loci, we crossed the line Flat-1
with our line HT1 (A,
Tl2, gp)
homozygous for the Hammarlund translocation (Table 2). (This
line was described in ref. 4). Analysis of 119 F2
plants revealed an obvious linkage of Tl2 with the translocation breakpoint
and the locus Gp residing near the breakpoint in
chromosome V
(here we use the linkage group numbers for desig-nation of the corresponding chromosomes). The major-ity (54 of 65) of the
structural heterozygotes, registered by pollen semi-sterility had the phenotype FT, that is, were hetero-zygotes tl2/+. The majority of fully fertile
plants (44 of 54) had wild-type tendrils. One can conclude that Tl2 resides near the translocation
breakpoint. In the line HT1 the longer interchange chromosome bears the allele gp near the T-point. Thirty-two
of 35 F2 plants
with the phenotype gp had
normal tendrils and only three had flat tendrils. These data provide evidence in favor of tl2 being located on a non-translocated
chromosome, either V or II.
We mentioned above that no linkage was observed between Tl2 and Cri, the latter being located on chromosome V immediately adjacent to the breakpoint. To explore the relationship between Tl2 and Cri further we examined F2 plants from the cross of our lines Cricytar (cri, a, r, Tl2) X Flat-1 (Cri, a, R, tl2/+), the F1 plants having a FT phenotype. Of 34 Cri plants in the F2, 18 had the wild-type phenotype and 16 had the FT phenotype. This ratio did not differ significantly from the ratio of these phenotypes among the cri F2 plants: 7 Tl : 5 FT. (c2= 0.105, P~0.75, calculated for a model implying equal proportion of FT plants in both classes Cri and cri). Because Tl2 and Cri do not appear to be linked and Cri is closer to the breakpoint than Gp (1, 4), Tl2 most probably resides on chromosome II. This hypothesis is supported by a decreased proportion of plants with the phenotype a (11 of 119, instead of 30 expected). Moreover, as discussed below, some of these 11 a plants were in fact trisomics. None of the a plants was semisterile or homozygous for the allele gp. These observations indicate that the a plants are structural homozygotes for the chromosomes of the standard karyotype.
 Self pollination of lines heterozygous for the Hammarlund translocation
is known to produce tertiary trisomics which have the karyotype represented by
two standard sets of chromosomes with an additional small interchange chromosome
(1, 4, 5). If the sporophytic
lethal tl2 resides
on the short arm of chromosome II above
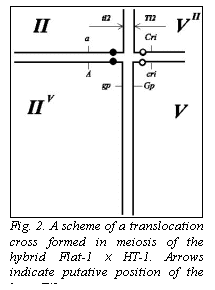
the breakpoint (Fig. 2), structural homozygotes for chromosome II will survive as trisomics if the
extra chromosome carries the wild-type allele
Tl2. Such a
trisomic would have the phenotype Gp a FT and fertile pollen. Of seven plants with this phenotype
four exhibited distinct external characteristics of trisomics.
An analysis of the karyotype in pollen mother cells revealed the presence
of seven bivalents plus a small univalent chromosome. The four plants with the
phenotype Gp a,
normal tendrils and fertile pollen could result from two cross-over events
between the loci A and Tl2. Thus, most probably, Tl2 resides in the short arm of chromosome II, although its location could be
determined more reliably in crosses involving plants with the normal karyotype.
Self pollination of lines heterozygous for the Hammarlund translocation
is known to produce tertiary trisomics which have the karyotype represented by
two standard sets of chromosomes with an additional small interchange chromosome
(1, 4, 5). If the sporophytic
lethal tl2 resides
on the short arm of chromosome II above
the breakpoint (Fig. 2), structural homozygotes for chromosome II will survive as trisomics if the
extra chromosome carries the wild-type allele
Tl2. Such a
trisomic would have the phenotype Gp a FT and fertile pollen. Of seven plants with this phenotype
four exhibited distinct external characteristics of trisomics.
An analysis of the karyotype in pollen mother cells revealed the presence
of seven bivalents plus a small univalent chromosome. The four plants with the
phenotype Gp a,
normal tendrils and fertile pollen could result from two cross-over events
between the loci A and Tl2. Thus, most probably, Tl2 resides in the short arm of chromosome II, although its location could be
determined more reliably in crosses involving plants with the normal karyotype.
The phenotype of heterozygote tl2/+ is very similar to that of tlw/+, although in tl2/+ the flat tendrils are somewhat wider and often are denticulate (Fig. 1). Moreover, the double heterozygote tl2/+, tlw/+ has a phenotype indistinguishable from that of the homozygote tlw/tlw. The only important difference is lethality of tl2 homozygotes. However, considering that the tl2 mutant was generated by g-radiation, the mutation might represent a small deletion that removes, in additon to tl2, an adjacent essential gene. According to its action, tl2 resembles a lethal mutation tlx in the locus tl described previously (2, 3).
Most legume species lack tendrils. It is therefore reasonable to suppose that conversion of the distal leaflets to tendrils appeared quite late in evolution, namely in the ancestors of the tribe Vicieae. This hypothesis is supported indirectly by viability of the null-mutation in the gene tl. However, loss-of-function mutation tl2, the effect of which in heterozygous state is almost indistinguishable from tlw/+, seems to arrest development in early stages of embryogenesis. We propose that the locus Tl appeared via duplication of the locus Tl2, which had some function important for viability. This hypothesis is supported by the observation that in both genomic regions the two Tl-type genes are linked to legumin genes and histone H1 genes (6). Sequencing of the genes Tl and Tl2 would provide an excellent test of our hypothesis.
Acknowledgement: This work was partly supported by the Russian State Program 'Russian Fund for Fundamental Research', grant No 99-04-49970.
1. Berdnikov, V.A., Gorel, F.L. and Temnykh. S.V. 1993. Pisum Genetics 25: 18-20.
2. Berdnikov, V.A., Gorel, F.L., Bogdanova, V.S., Kosterin, O.E., Trusov, Y.A. and Rozov. S.M. 1999. Genet. Res. 73: 93-109.
3. Gorel, F.L., Berdnikov, V.A. and Temnykh. S.V. 1994. Pisum Genetics 26: 16-17.
4. Gorel, F.L., Kosterin, O.E. and Berdnikov. V.A. 1999. Pisum Genetics 31:5-8.
5. Pellew, C. 1940. J. Genet. 39: 363-390.
6. Weeden, N.F., Ellis, T.H.N., Timmerman-Vaughan, G.M., Swiecicki, W.K., Rozov, S.M. and Berdnikov. V.A. 1998. Pisum Genetics 30: 1-4.