Pisum
Genetics Volume 27 1995 Research Reports pages 12-13
Enod2 exhibits linkage with a (linkage group IA) and Fum (linkage
group II)
Paruvangada, V. G., Weeden, N.F., Department of Horticultural Sciences
Cargnoni, T. and Yu, J. NYSAES, Cornell University
Geneva,
NY 14456, USA
Gorel',
F. Institute
of Cytology and Genetics
Novosibirsk
630090, Russia
Frew, T., Timmerman-Vaughan, G.M. and Crop and Foods Research
McCallum, J. Private Bag 4704, Christchurch, New Zealand
The gene a traditionally has been placed on the
upper portion of linkage group I (2, 11). However, recent
work by Kosterin and others (4, 5) has indicated linkage group I may actually
represent two linkage groups, tentatively designated IA and IB. Linkage group
IB covers 70 to 150 cM, stretching from the d — sym2 region to af— i and
perhaps somewhat beyond (3, 11). Linkage
group IA includes Aat-p, His2-6, a, lf, and His7, but is
relatively short (50-100 cM) and thus a good candidate for synteny with
one of the other linkage groups described for pea. Here we present results identifying linkage between a nodulin gene, Enod2,
and markers on both linkage group IA and II, suggesting that these
two linkage groups may be on the same chromosome.
The
Enod2 clone was generously supplied by T. Bisseling (Agricultural University,
Wageningen). Another DNA clone, CD40 (=Lg-J),
was mapped by Ellis et al. (3). The Enod2 gene codes a hydroxyproline-rich protein expressed early in nodule
development (10); however, the
specific function of the protein has yet to be determined. The probe was radiolabelled and used for Southern hybridization
analysis as described in Polans et al. (7). Two populations were
examined. A set of F8 recombinant inbred lines (RILs) from the cross
JI1794 x Slow and a second set of 60 F6
RILs from the cross Sparkle x WL1238. In both populations polymorphism
was observed using the restriction endonuclease, EcoRI.
In both populations, Enod2 gave approximately a
1:1 segregation ratio (18:25 in JI1794 x Slow
and 22:20 in Sparkle x WL1238), as did most of the several hundred other
markers scored (data not presented), indicating that
there were no major differences in karyotype between each pair
of parents. In the JI1794 x Slow population, joint segregation analysis for Enod2 and
Fum gave 11 recombinants out of the 42 lines scored for both markers and
the analysis for Enod2 and His2-6 gave 14 recombinants out
of 42 lines. Using the equation r = R/[2-2R]
(8), which relates recombination frequency (r) to the proportion of RILs in
which recombination has occurred
between two loci (R), we obtained a recombination frequency between Enod2
and Fum of 18% and one between Enod2 and His2-6 of
25%. Although these linkages are not
particularly tight, we have located several other RFLP and RAPD markers in the
region, generating a much more saturated map and confirming the linkages (Fig.
1).
In the Sparkle x
WL1238 RIL population Enod2 again showed linkage with markers on linkage group IA : a - 20% - B319 - 9% - Enod2.
However, in this population we did not observe linkage between Enod2 and the markers on linkage group II
including wb, k, and Pgm-p. These latter markers are
over 20 cM from Fum on the side opposite Enod2, and the relatively
few RILs in the population may have prevented our observing significant
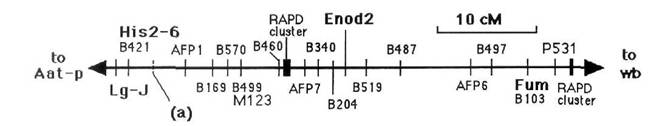
Fig. 1. Linkage group for the JI1794 x Slow population that contains Enod2
and markers on both linkage group IA (His2-6, Lg-J) and on linkage
group II (Fum). These standard markers are given in bold print, two cDNA markers (M123 and P531) are given in
medium sized print, and RAPDs are given in small
print. Two RAPD clusters containing 5 or more RAPDs each, are also mapped. The approximate position of gene a
is shown for convenience, although it did not segregate in this population.
deviations from
random assortment Unfortunately, Fum was not segregating in this
population, nor have we identified many DNA
markers in this region of the genome in this set of RILs. Enod2 did not display significant linkage with any other
region of the genome in either of the RIL populations.
We
conclude that Enod2 is located 25 to 30 cM from a on linkage
group IA. Our results support the suggestion
by Kosterin that a and d are located on different linkage groups,
because Enod2 did not display linkage with any marker on linkage
group IB. Rather, it appears that group
IA is syntenic with Fum, forcing the joining of group IA with linkage
group II. On cytogenetic grounds such a merger would be acceptable because
group IA is assigned to chromosome 6, but appears to constitute only a
portion of this chromosome, whereas group II has
not been assigned to a chromosome (9).
The location of Enod2 does not correspond to that of any
previously mapped sym or Enod gene (6). Two sym genes (sym14 and that from JI1794,
tentatively designated Sym22) are near Fum,
but appear to be on the side of Fum opposite to Enod2 (S.
Temnykh, T.A. LaRue, and N.F. Weeden,
unpublished). Another non-nodulating gene, Twt, lies very near a (1). Thus, Enod2 must be a new symbiosis gene that can not be
paired with any of the known symbiosis mutants.
1.
Berdnikov, V.A.,
Gorel', F.L. and Temnykh, S.V. 1994. Pisum Genet. 26:9-10.
2.
Blixt,S. 1972. Agri Hort.
Genet. 30:1-293.
3.
Ellis, T.H.N.,
Hellens, R.P., Turner, L., Lee, C, Domoney, C. and Welham, T.
1993. Pisum Genet. 25:5-12.
4.
Kosterin, O.E. 1993.
Pisum Genet. 25:23-26.
5.
Kosterin, O.E. and
Rozov, S.M. 1993. Pisum Genet. 25:27-31.
6.
LaRue, T.A. and
Weeden, N.F. 1992. Pisum Genet. 24:5-12.
7.
Polans, N.O., Weeden,
N.F. and Thompson, W.F. 1985. Proc. Natl. Acad. Sci.
USA 82:5083-5087.
8.
Taylor, B.A. 1978. In:
Origins of Inbred Mice. H.C. Morse (ed.). Academic Press,
New York, pp. 432-438.
9.
Temnykh, S.V. and
Weeden, N.F. 1993. Pisum Genet. 25:1-3.
10.
van der Weil, C.,
Sheres, B., Franzen, H., van Lierop, M.-J., van Lammeren, A., van
Kammen, A. and Bisseling, T. 1990. EMBOJ. 9:1-7.
11.
Weeden, N.F.,
Ambrose, M. and Swiecicki, W.K. 1991. Pisum Genet. 23:cover.
