|
Pisum Genetics |
Volume 26 |
1994 |
Research Reports |
pages 9-10 |
Gene Twt (twisted tendrils) decoupled from the translocation breakpoint became a convenient dominant marker on chromosome 1.
|
Berdnikov, V.A., Gorel, F.L. and Temnykh, S.V. |
Institute of Cytology and Genetics Novosibirsk 630090, Russia |
Earlier we described a new dominant mutation Twt (twisted tendrils) (1) tightly linked to the break point of a translocation which arose simultaneously with the mutation. The translocation involves chromosome 1 and an unidentified chromosome. Among 106 plants obtained by setting of a structural heterozygote (1), all 43 plants with normal tendrils had fertile pollen, while all 52 plants with phenotype Twt had semisterile pollen. We concluded that gene Twt must be located in close proximity (less than 1 cM) to the break point. In the progeny of a cross between the structural heterozygote and plants possessing a normal karyotype, the tertiary trisomic (2n + 1-X) was identified (1). The additional chromosome, 1-X, carried a fragment of chromosome 1, with genes A (anthocyanin), His(2-6) (encoding subtypes 2-6 of histone H1), and Twt. Trisomic plants had phenotype Twt and fertile pollen, but were inferior to their 2n siblings in vigour and fertility. Selfing of these trisomic plants should have produced the following karyotypic classes: trisomics, structural heterozygotes, homozygotes for reciprocal translocations, or homozygotes for normal karyotype; the latter class of plants should have phenotype twt and the others phenotype Twt. However, one fertile and vigorous plant with phenotype Twt was detected in the progeny of a trisomic. It gave 249 seeds in contrast to the 5-10 seeds ordinarily produced by trisomics. This exceptional plant turned out to be heterozygous for loci a and His(2-6) [phenotype 1021/1133, for designation of H1 haplotypes see (1)] and homozygous for gene His7 (allele 2) encoding H1 subtype 7 (2).
We examined 216 individual progeny from this plant. All progeny proved to be fully fertile and showed segregation for phenotypes A : a and Twt : twt. Chromosome analysis of pollen mother cells of representatives of all the phenotypic classes showed the presence of the common, seven bivalent configuration at meiotic metaphase I. The pattern of phenotypic segregation (Table 1) allowed us to consider gene Twt as a dominant factor linked to genes a and His(2-6). The calculated recombination fractions for gene pairs His(2-6) – a and His(2-6) – Twt are 5.5 ± 1.6 and 12.5 ± 2.4, respectively.
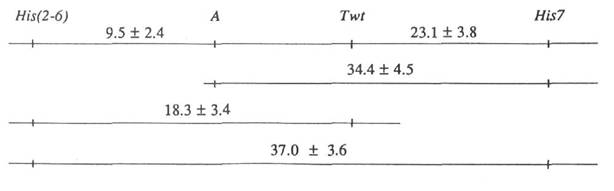
One of these plants with phenotype His(2-6)1133 , a, Twt, His72 was crossed with the line WL1255 with genotype His(2-6)1221, A, twt, His73. Among five F1 plants, three had phenotype Twt and two had phenotype twt indicating the parental plant was heterozygous at this locus. The three Twt plants were selfed and the resulting F2 scored for segregating linkage group I markers. The combined data are presented in Table 2. Recombination fractions ± SE were obtained using the method of maximum likelihood with the aid of the computer program CROS developed by S.M. Rozov. The results can be presented as the following segment of the genetic map for linkage group 1:

(The distance between genes a and Twt appeared to be zero due to their repulsion phase and the absence of double cross-overs with phenotype a twt; it is impossible to estimate the actual distance between closely linked loci in such a cross). This map corresponds well to the recently reported (2) normal order of genes His(2-6), A , and His7.
Table 1. Joint segregation of loci His(2-6), a, and Twt in the progeny of the exceptional trisomic descendant with normal fertility and karyotype. (The total number of plants is 216).
|
Anthocyanin |
a |
A |
||||
|
His(2-6) |
1021 |
1021/1133 |
1133 |
1021 |
1021/1133 |
1133 |
|
Twt |
36 |
7 |
0 |
4 |
92 |
8 |
|
twt |
2 |
0 |
0 |
0 |
17 |
50 |
Table 2. Joint segregation of loci Twt, A, His(2-6), and His7. (Total number of plants is 159).
|
His(2-6) |
1133 |
1133/1221 |
1221 |
||||
|
Anthocyanin |
A |
a |
A |
a |
A |
a |
|
|
His7 |
Twisted tendrils |
|
|
|
|
|
|
|
|
Twt |
1 |
2 |
12 |
0 |
4 |
0 |
|
3 |
|
|
|
|
|
|
|
|
|
twt |
0 |
0 |
11 |
0 |
13 |
0 |
|
|
Twt |
5 |
14 |
36 |
2 |
7 |
0 |
|
2/3 |
|
|
|
|
|
|
|
|
|
twt |
0 |
0 |
1 |
0 |
9 |
0 |
|
|
Twt |
2 |
11 |
21 |
1 |
4 |
1 |
|
2 |
|
|
|
|
|
|
|
|
|
twt |
0 |
0 |
1 |
0 |
1 |
0 |
The cause of the observed separation of gene Twt from the translocation breakpoint is not clear, it might be crossing over or some reverse translocation. Anyway, this gene became a convenient dominant marker on chromosome 1. Moreover, the line appears to possess a normal karyotype.
Gorel, F.L., Temnykh, S.V., Lebedeva,
I.P. and Berdnikov, V.A. 1992. Pisum Genetics
24:48-51.
Kosterin, O.E. 1992. Pisum Genetics 24:56-59.