Wolko, B. and N. F. Weeden NYS
Agricultural Experiment Station,
Geneva, NY 14456
USA
Enzymes
catalyzing the reaction:
NADH + acceptor = NAD + reduced
acceptor can be visualized
after horizontal starch gel
electrophoresis by using an assay consisting of 100 ml 0.1 M Tris HC1 pH
8.5, 40 mg NADH, 40 mg MTT and
1 mg 2,6 dichlorophenol indophenol. At least four NADH
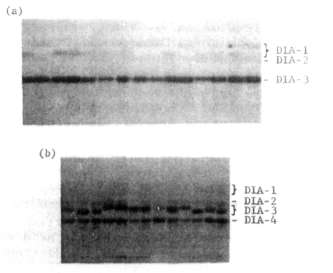
diaphorases (DIA) isozymes can be resolved in pea leaf extracts, and we
have found variation in the most anodal isozyme DIA-1 and the
most intensely staining isozyme DIA-3 (Fig. 1). The DIA-1 polymorphism is best
resolved using the pH 6.5 histidine buffer system of Cardy et
al. (1), whereas the DIA-3 variation is more clearly observed on a Tris borate-EDTA system (2). In a survey of a wide sample of Pisum germplasm, we identified
at least 2 common variants for DIA-1 the more anodal of which we designated "a" and the other "b". We demonstrate here
that the variation in DIA-1 phenotype shows monogenic inheritance, being encoded by a locus
that exhibits linkage with
markers near M on chromosome 3.
Marker lines fixed for DIA-la were
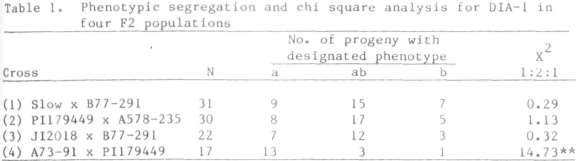
crossed with lines fixed for DIA-lb, and the resulting hybrids selfed to form F2 progenies. Segregation for DIA-1
phenotype was observed in each of the four F2 progenies analyzed (Table
1). In three of the four progenies the DIA-1 variants behaved as codominant alleles at a single segregating
locus, which we designated Dia-1. The fourth progeny derived from the cross A73-91 x PI 179449 gave all three of the expected phenotypes but the
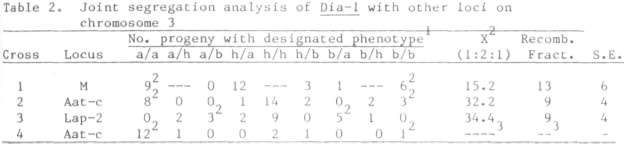
relative number of these was significantly different from the expected 1:2:1 ratio. Joint segregation analysis of the loci segregation in
these progenies indicated linkage between Dia-1 and loci near M and chromosome
3 (Table II).
Previous results indicate that
Aat-c is about 15 map units from M and Lap-2 (3).
Comparative map distances and the lack of linkage between Dia-1 and
Acp-3 or St (results not
presented) suggest that
Dia-1 is located about midway between Lap-2 and Aat-p on the
distal side of M from the
centromere. The availability
of two common alleles at
Dia-1 should make this locus very useful in further mapping studies.
This work was supported in part
by a grant from the International Board of Plant Genetics
Resources (Grant #86/102).