LINKAGE DETERMINATIONS FOR SEVERAL IS02YMIC LOCI IN PISUM
Weeden, N, F. and G. A. Marx NYS Agricultural Experiment Station
Geneva, NY, USA
A number or isozyme systems have been described in Pisum
(1,2,5,6,9), but only one locus, that specifying a leucine
aminopeptidase, has been mapped on the pea genome (1). We have
initiated a program aimed at producing an isozyme linkage map for the
entire Pisum genome. This report presents data comfirming the position
of the gene coding Lap-1 and describes approximate map positions of two
other isozyme loci.
The parents and three F2 progenies were grown in the greenhouse.
The two parental lines for progeny C282-23 1 contained the following
markers: Female (WL 1466): A, D, I, b am-2. Pl, R
Male: a, i, mo, En, b, er., z, r
In aadition, the female parent exhibited a Lap-1 (fast), 6 pgd-2 (fast)
phenotype while the male possessed the "slow" allele at both of these
loci. For progenies C282-232 and C282-233 the parents possessed
markers: Female: A, Dco , I, k, st, f, b, le, fs, wlo, tl, R
Male: a, i_, k, gp, cov., te, Fs, p, R
Progeny C282-232 segregated for allozymes at Acp-1 and Acp-3, in
addition to the other known markers.
Isozyme phenotypes were determined by starch gel electrophoresis,
performed as described by Gottlieb (4). Two buffer systems were used in
the preparation of the gels, a tris-citrate/lithium borate system at pH
8.1 (7) and an N-(3~aminopropy1)-morpholine/citric acid buffer at pH 6.1
(3). Assay mixtures were slight modifications of those described in
Shaw and Prasad (8). Linkage between loci was analyzed by comparing
observed and expected joint segregation ratios in pairwise combinations.
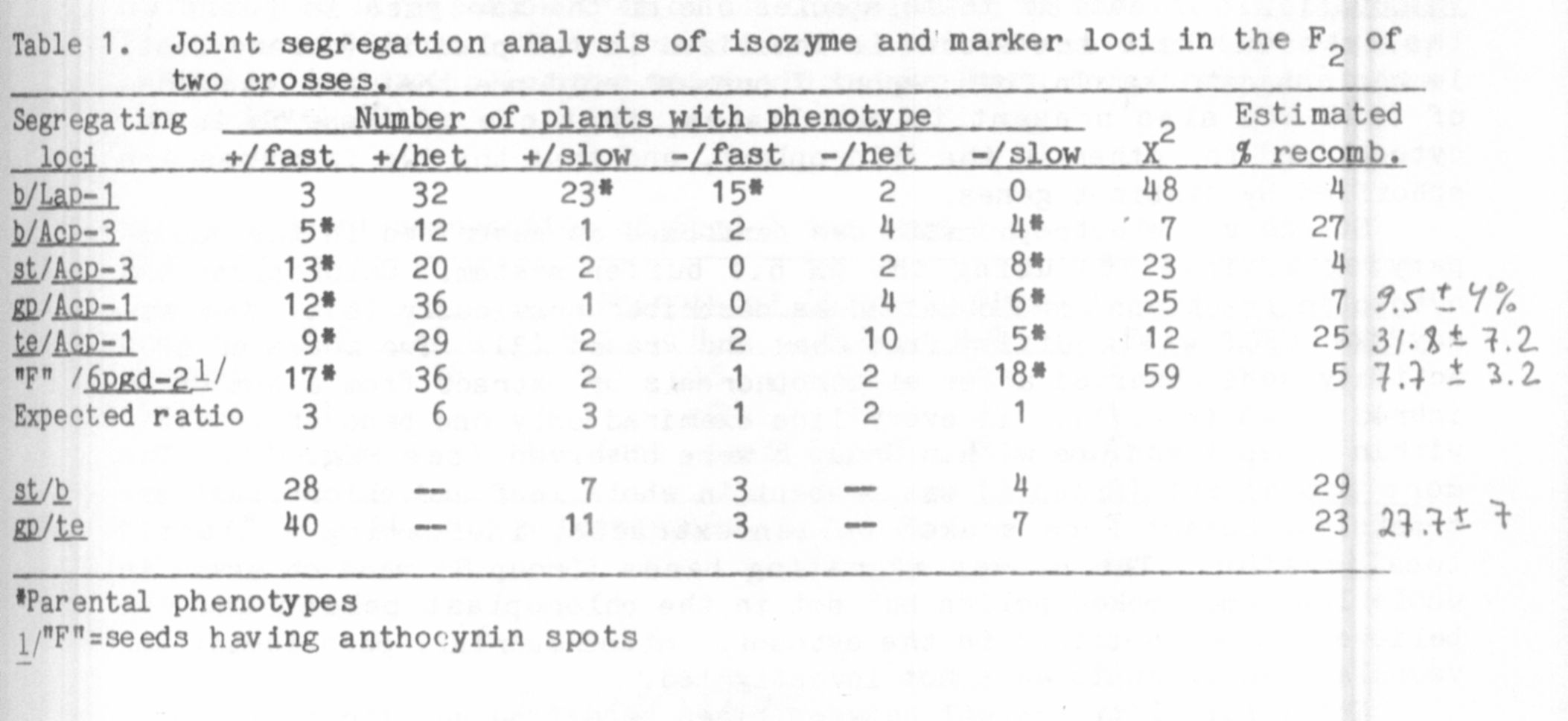
Tight linkage was observed between the b locus on chromosome 3 and
Lap-1 (Table 1), thus corroborating a previous report (1). Our data
gave a recombinant frequency between the two loci of about 4%. A second
isozyme locus, Acp-3. can also be placed on chromosome 3- Acp-3 was
found to give 9% recombinants with respect to st and 32% with b, while
24% recombination was observed between st and b (Table 1). Thus, the
linkage order appears to be Acp-3—st:—b.
A second acid phosphatase locu3, Acp-1 , assorted independently of
Acp-3. but showed non-random joint segregation ratios with two markers,
gp and te on chromosome 5. Recombination between gp and Acp-1 was 7%
and that between te and Acp-1 was 25% (Table 1). The estimated
recombination between the two marker loci was 23%, suggesting that Acp-1
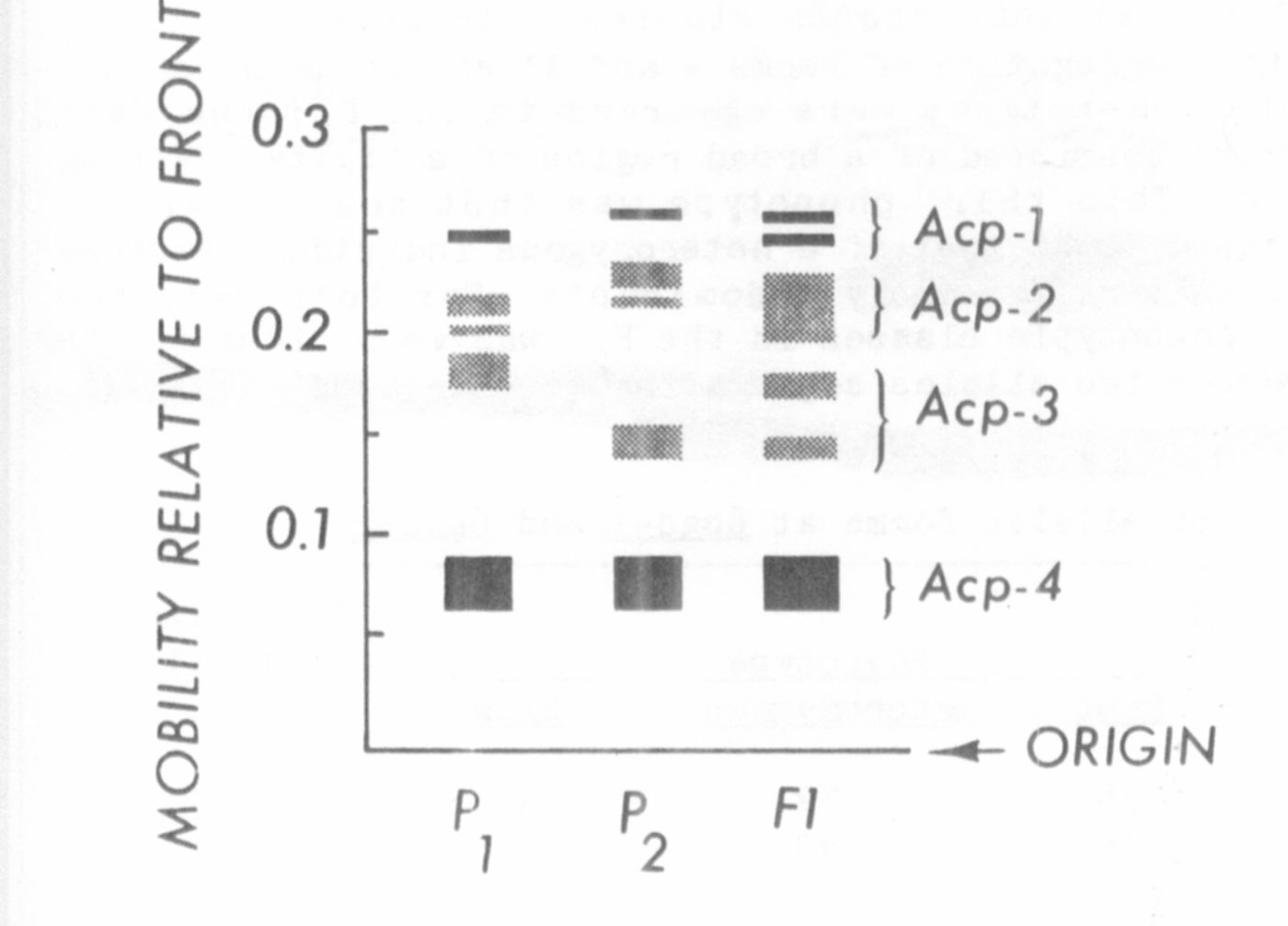
is located distal to gp, i.e. toward cp. The relative mobilities of the
acid phosphatase isozymes are shown in Fig. 1.