ELECTROPHORETIC STUDY OF SEVERAL ENZYME SYSTEMS OF THE GENUS P1 SUM WITH REFERENCE TO ITS
TAXONOMY
Przybylska, J., H. Parzysz, Institute of Plant Genetics, Poznan, Poland
and S. Blixt Weibullsholm Plant Breeding Institute, Landskrona, Sweden
In an investigation initiated recently, iso-enzyme analysis was used to
explore the taxonomic relationship among diverse forms of Pisum. This report
presents data on distribution of electrophoretic phenotypes of several
enzyme systems: leucine aminopeptidase (LAP), glutamate oxaloacetate trans-
aminase (GOT), and peroxidases (PX) .
A total of 108 lines were examined, including P_. elatius (4 lines),
humi le (3 lines), P. sativum (84 lines), P_. abyssinicum (12 lines), and
P_. fulvum (5 lines). Most of the lines originated from the Weibullsholm
Collection. Cotyledons of mature seeds (LAP), as well as leaves (GOT) and
roots (PX) of 2-3-week-old seedlings were analyzed. An electrophoretic analysis
of LAP and GOT systems was performed in starch gel, and that of the PX system
in polyacrylamide gel.
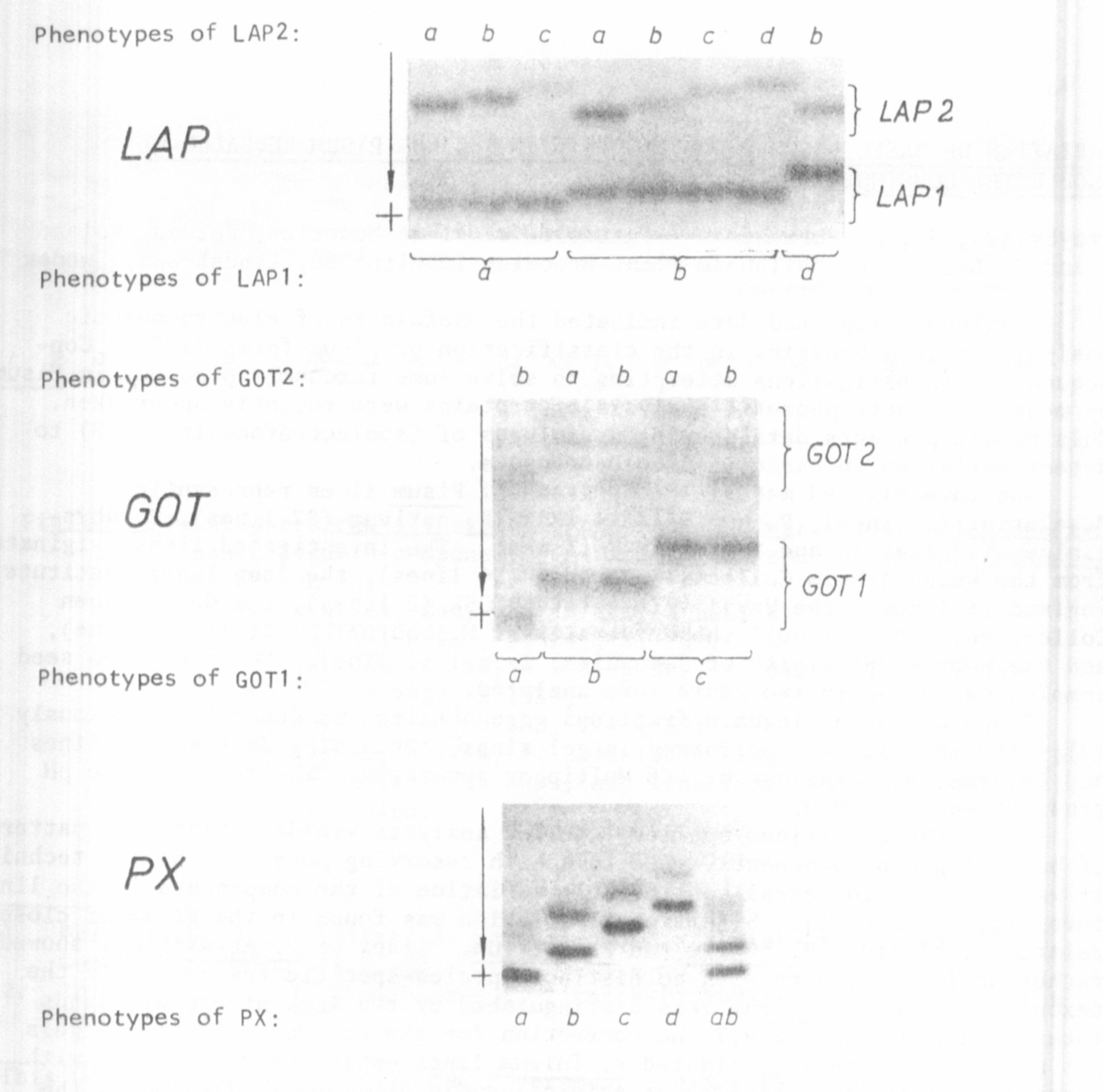
Zymograms showed the following variation in electrophoretic phenotypes:
LAP - two zones of activity exhibiting independent variation, LAP1 and LAP2
with 3 and 4 phenotypes, respectively; GOT - two zones of activity exhibiting
independent variation, G0T1 and G0T2 with 3 and 2 phenotypes, respectively;
PX - one variant zone with 5 phenotypes (Fig. 1). In the 5 enzyme zones,
representing probably at least 5 loci, 17 phenotypes were observed in total.
In considering the distribution of the different electrophoretic patterns
that were found, two kinds of characters were taken into account: 1) Electro-
phoretic phenotypes of particular enzyme zones - assumed to be products of
allelic genes at single loci, and 2) Combinations of electrophoretic phenotypes
of different enzyme zones, i.e. combinations of phenotypes assumed to be
controlled by various loci, named here Composite Electrophoretic Phenotypes
(CEPHs).
The distribution of the electrophoretic phenotypes tentatively attributed
to allelic genes at single loci, considered in terms of presence or absence
in particular taxons, showed no species-specific characters. It can, however,
be expected that when the investigated material is extended, differences
between the examined taxons may be reflected in various frequencies of certain
electrophoretic phenotypes.
As regards the distribution of the 29 CEPHs that were distinguished,
the variation found in P. sativum (22 CEPHs) partly overlapped that found
in P. elatius and P. humile. All the investigated P. abyssinicum lines showed
the same CEPH, not observed in other taxons. Nor were three CEPHs found in
P. fulvum observed in other forms. Grouping of Pisum forms on the basis of
this enzyme analysis was similar to the classification based on electrophoretic
patterns of seed albumins (3) and globulins (4). P. fulvum and P. abyssinicum
appear to be different taxons, as distinguished from the group which showed
greater similarity, viz. P. elatius, P. humile, and P. sativum.
Polymorphism of aminopeptidases (1,5), usually referred to as "leucine
aminopeptidase", and that of peroxidases (2) in Pisum was reported earlier.
The wide range of genetic variation presently analyzed revealed additional
polymorphism of leucine aminopeptidase.